Are you sure you want to delete this access key?




Python package for concise, transparent, and accurate predictive modeling. All sklearn-compatible and easy to use.
docs • imodels overview • demo notebooks

Modern machine-learning models are increasingly complex, often making them difficult to interpret. This package provides a simple interface for fitting and using state-of-the-art interpretable models, all compatible with scikit-learn. These models can often replace black-box models (e.g. random forests) with simpler models (e.g. rule lists) while improving interpretability and computational efficiency, all without sacrificing predictive accuracy! Simply import a classifier or regressor and use the fit and predict methods, same as standard scikit-learn models.
from imodels import BoostedRulesClassifier, BayesianRuleListClassifier, GreedyRuleListClassifier, SkopeRulesClassifier # see more models below
from imodels import SLIMRegressor, RuleFitRegressor
model = BoostedRulesClassifier() # initialize a model
model.fit(X_train, y_train) # fit model
preds = model.predict(X_test) # discrete predictions: shape is (n_test, 1)
preds_proba = model.predict_proba(X_test) # predicted probabilities: shape is (n_test, n_classes)
print(model) # print the rule-based model
-----------------------------
# the model consists of the following 3 rules
# if X1 > 5: then 80.5% risk
# else if X2 > 5: then 40% risk
# else: 10% risk
Install with pip install imodels (see here for help).
| Model | Reference | Description |
|---|---|---|
| Rulefit rule set | 🗂️, 🔗, 📄 | Fits a sparse linear model on rules extracted from decision trees |
| Skope rule set | 🗂️, 🔗 | Extracts rules from gradient-boosted trees, deduplicates them, then linearly combines them based on their OOB precision |
| Boosted rule set | 🗂️, 🔗, 📄 | Sequentially fits a set of rules with Adaboost |
| Slipper rule set | 🗂️, ㅤㅤ 📄 | Sequentially learns a set of rules with SLIPPER |
| Bayesian rule set | 🗂️, 🔗, 📄 | Finds concise rule set with Bayesian sampling (slow) |
| Optimal rule list | 🗂️, 🔗, 📄 | Fits rule list using global optimization for sparsity (CORELS) |
| Bayesian rule list | 🗂️, 🔗, 📄 | Fits compact rule list distribution with Bayesian sampling (slow) |
| Greedy rule list | 🗂️, 🔗 | Uses CART to fit a list (only a single path), rather than a tree |
| OneR rule list | 🗂️, ㅤㅤ📄 | Fits rule list restricted to only one feature |
| Optimal rule tree | 🗂️, 🔗, 📄 | Fits succinct tree using global optimization for sparsity (GOSDT) |
| Greedy rule tree | 🗂️, 🔗, 📄 | Greedily fits tree using CART |
| C4.5 rule tree | 🗂️, 🔗, 📄 | Greedily fits tree using C4.5 |
| Iterative random forest |
🗂️, 🔗, 📄 | Repeatedly fit random forest, giving features with high importance a higher chance of being selected |
| Sparse integer linear model |
🗂️, ㅤㅤ📄 | Sparse linear model with integer coefficients |
| Sapling Sums | 🗂️, ㅤㅤ📄 | Sum of small trees with very few total rules (SAPS) |
| Shrunk trees wrapper |
🗂️, ㅤㅤ📄 | Use regularization to improve trees |
| Distillation wrapper |
🗂️, ㅤㅤ📄 | Train a black-box model, then distill it into an interpretable model |
| More models | ⌛ | (Coming soon!) Lightweight Rule Induction, MLRules, ... |
Docs 🗂️, Reference code implementation 🔗, Research paper 📄
| Discretizer | Reference | Description |
|---|---|---|
| MDLP | 🗂️, 🔗, 📄 | Discretize using entropy minimization heuristic |
| Simple | 🗂️, 🔗 | Simple KBins discretization |
| Random Forest | 🗂️ | Discretize into bins based on random forest split popularity |
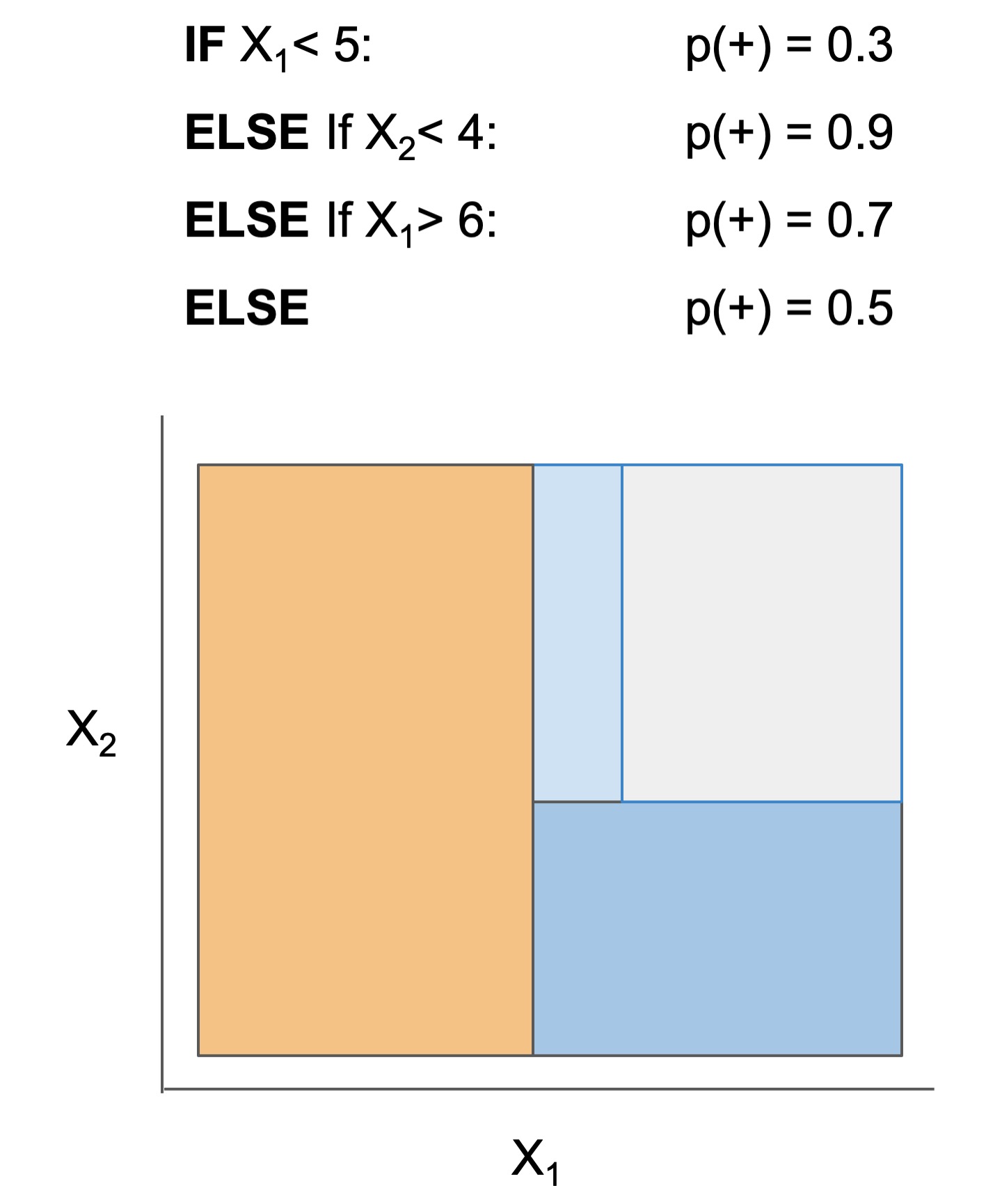
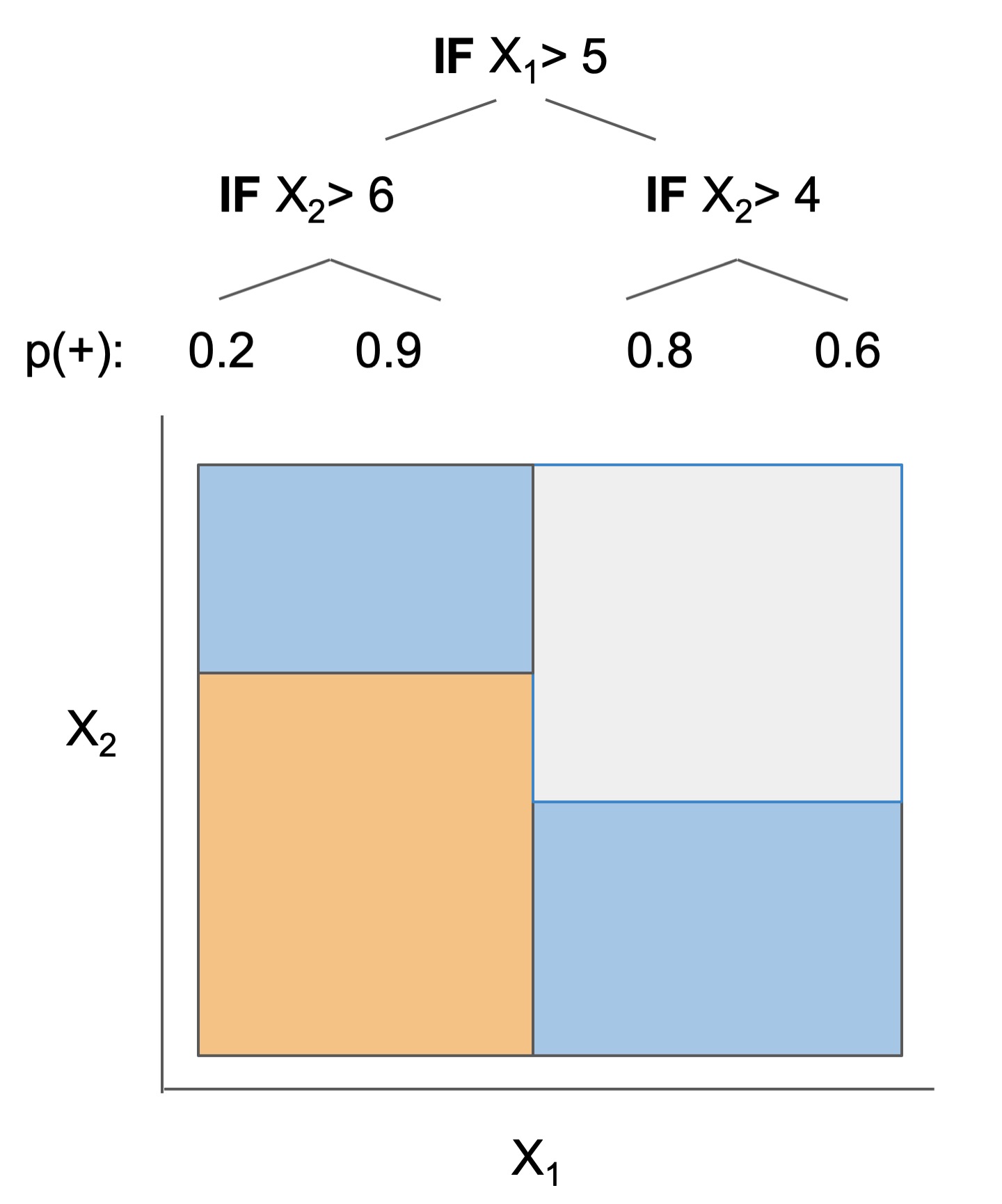
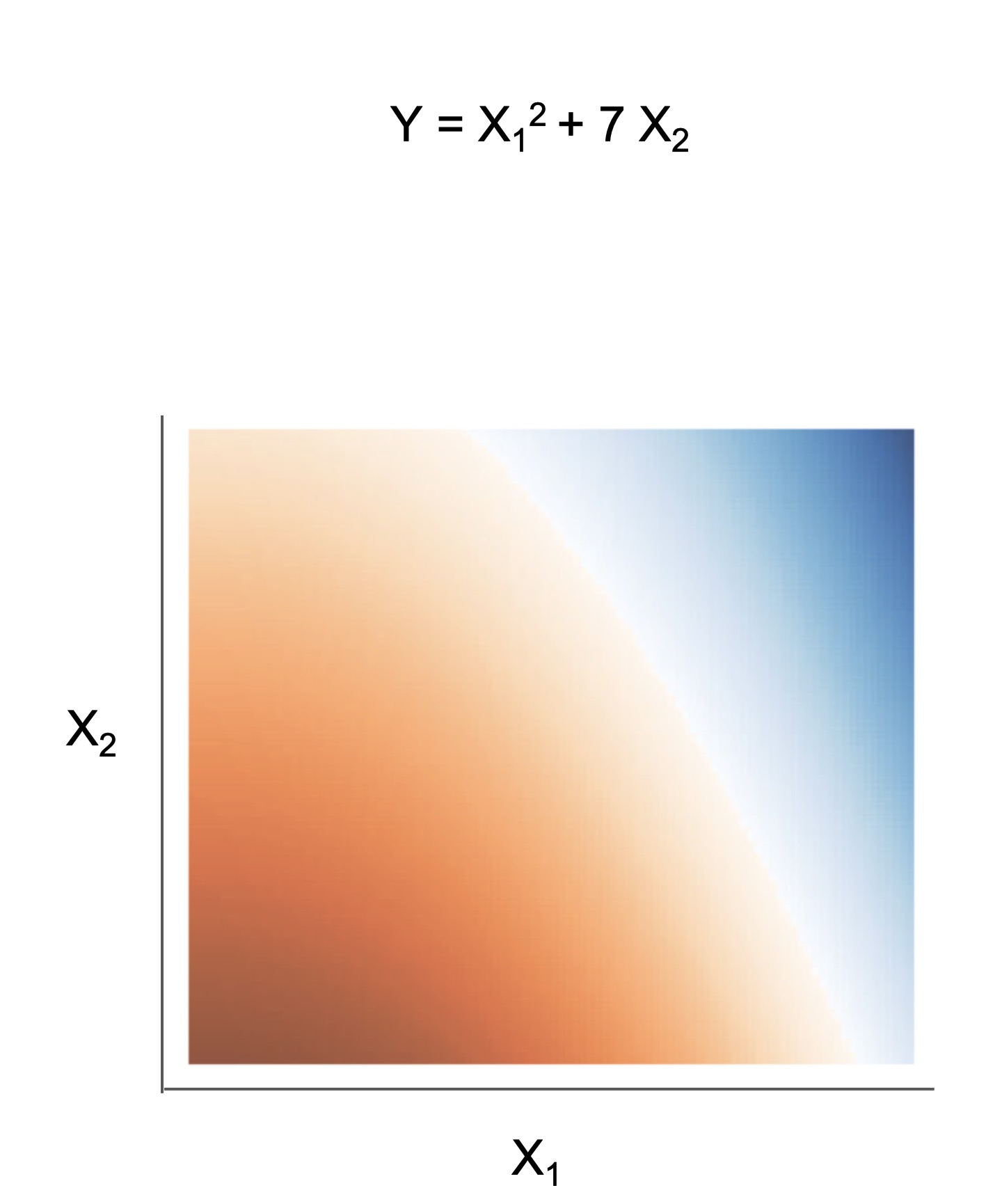
The final form of the above models takes one of the following forms, which aim to be simultaneously simple to understand and highly predictive:
| Rule set | Rule list | Rule tree | Algebraic models |
|---|---|---|---|
 |
 |
 |
 |
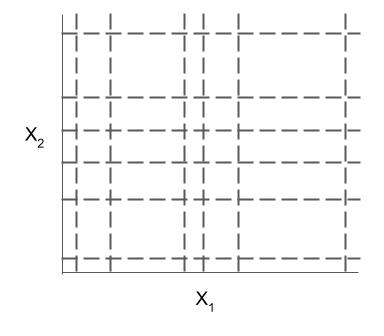
Different models and algorithms vary not only in their final form but also in different choices made during modeling. In particular, many models differ in the 3 steps given by the table below.
See the docs for individual models for futher descriptions.
| Rule candidate generation | Rule selection | Rule postprocessing |
|---|---|---|
 |
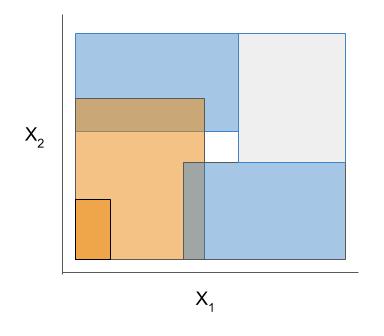 |
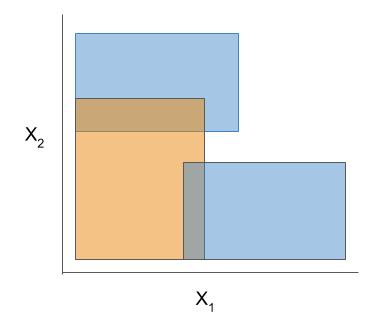 |
The code here contains many useful and customizable functions for rule-based learning in the util folder. This includes functions / classes for rule deduplication, rule screening, and converting between trees, rulesets, and neural networks.
Demos are contained in the notebooks folder.
imodels for deriving a clinical decision rule
Different models support different machine-learning tasks. Current support for different models is given below (each of these models can be imported directly from imodels (e.g. from imodels import RuleFitClassifier):
| Model | Binary classification | Regression | Notes |
|---|---|---|---|
| Rulefit rule set | RuleFitClassifier | RuleFitRegressor | |
| Skope rule set | SkopeRulesClassifier | ||
| Boosted rule set | BoostedRulesClassifier | ||
| SLIPPER rule set | SlipperClassifier | ||
| Bayesian rule set | BayesianRuleSetClassifier | Fails for large problems | |
| Optimal rule list (CORELS) | OptimalRuleListClassifier | Requires corels, fails for large problems | |
| Bayesian rule list | BayesianRuleListClassifier | ||
| Greedy rule list | GreedyRuleListClassifier | ||
| OneR rule list | OneRClassifier | ||
| Optimal rule tree (GOSDT) | OptimalTreeClassifier | Requires gosdt, fails for large problems | |
| Greedy rule tree (CART) | GreedyTreeClassifier | GreedyTreeRegressor | |
| C4.5 rule tree | C45TreeClassifier | ||
| Iterative random forest | IRFClassifier | Requires irf | |
| Sparse integer linear model | SLIMClassifier | SLIMRegressor | Requires extra dependencies for speed |
| Sapling Sums (SAPS) | SaplingSumClassifier | SaplingSumRegressor | |
| Shrunk trees | ShrunkTreeClassifierCV | ShrunkTreeRegressorCV | Wraps any sklearn tree-based model |
| Distillation | DistilledRegressor | Wraps any sklearn-compatible models |
If it's useful for you, please star/cite the package, and make sure to give authors of original methods / base implementations credit:
@software{
imodels2021,
title = {{imodels: a python package for fitting interpretable models}},
journal = {Journal of Open Source Software}
publisher = {The Open Journal},
year = {2021},
author = {Singh, Chandan and Nasseri, Keyan and Tan, Yan Shuo and Tang, Tiffany and Yu, Bin},
volume = {6},
number = {61},
pages = {3192},
doi = {10.21105/joss.03192},
url = {https://doi.org/10.21105/joss.03192},
}
Press p or to see the previous file or, n or to see the next file
Are you sure you want to delete this access key?
Are you sure you want to delete this access key?
Are you sure you want to delete this access key?
Are you sure you want to delete this access key?